Speaker
Description
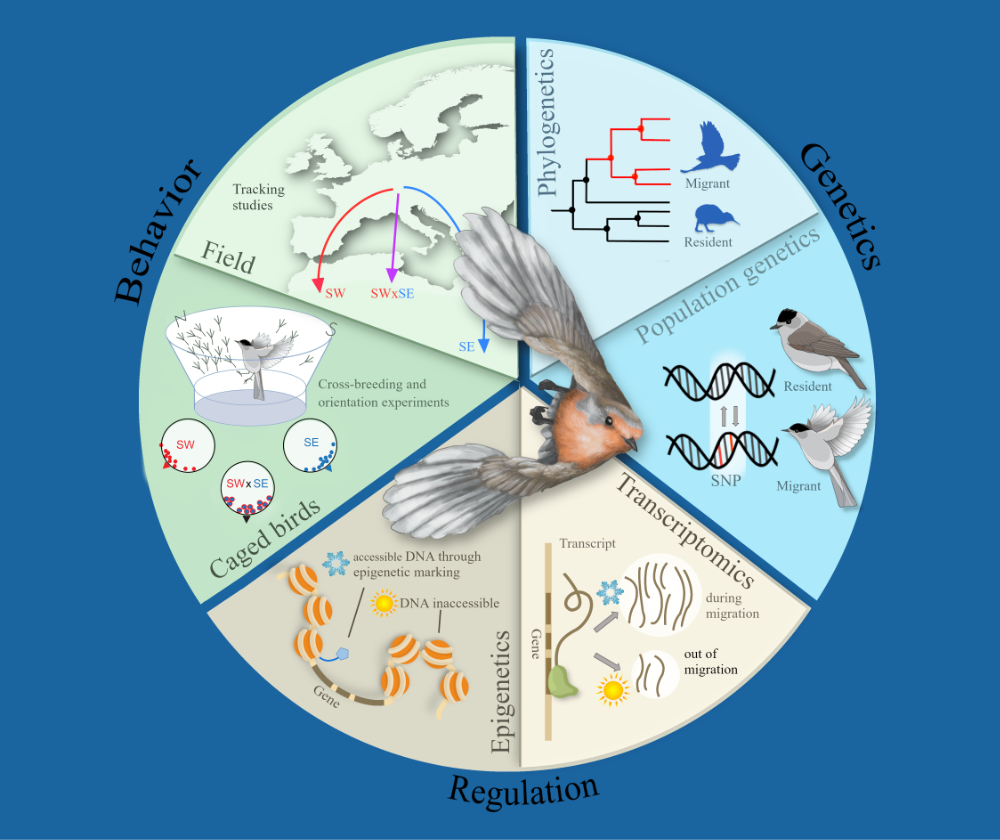
Avian migration is an ecologically and evolutionarily important behaviour. Classic breeding experiments show that the difference in migratory propensity between migrant and resident populations has genetic bases. More recent population genomic studies indicate that differential selection on gene regulation instead of protein variants may be responsible for this heritable behavioural variation. However, the molecular regulation and its cell type-specificity underlying this behavioural variation between populations are still unknown. To address differential molecular regulation between populations with different propensity to migrate at the single cell level, I perform single nucleus Assay for Transposase-Accessible Chromatin sequencing (snATAC-seq) to study chromatin accessibility in blackcaps (Sylvia atricapilla) from a short distance migratory and one resident population during migratory and a control condition outside the migratory season. I specifically focus on the hypothalamus, which is a brain structure with conserved anatomy, cell types and functions regulating adaptive physiology and behaviour relevant for seasonal migration. I will present preliminary results of three analyses: (1) motif enrichment - asking what cis-regulatory elements in what cell types are differentially accessible between populations and seasons; (2) differential transcription factor footprinting - addressing what transcription factors are differentially bound to cis-elements between populations and seasons; and (3) characterisation of cis-co-accessibility networks describing sets of co-regulated cis elements. This study highlights the relevance of heterogeneity on the single-cell level and interaction of multiple regulatory elements in studying molecular mechanisms of behavioural variation.